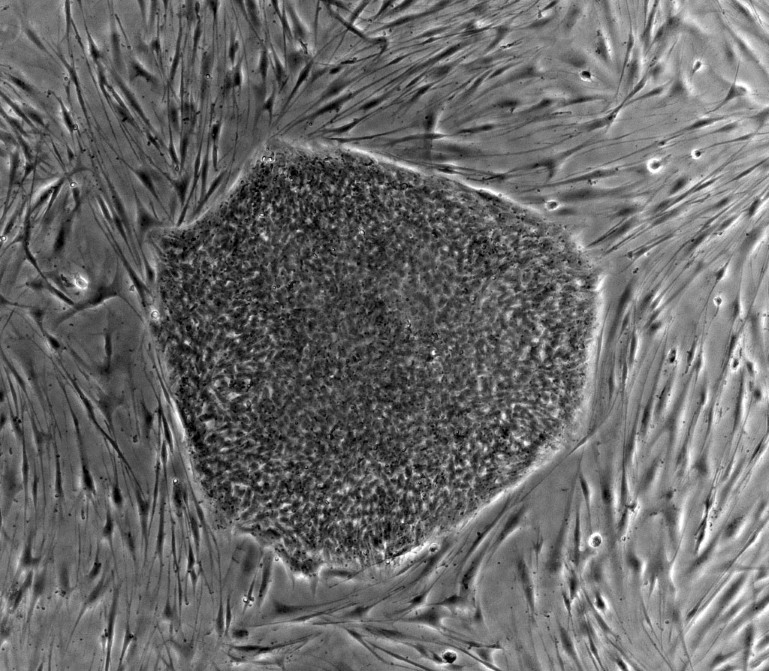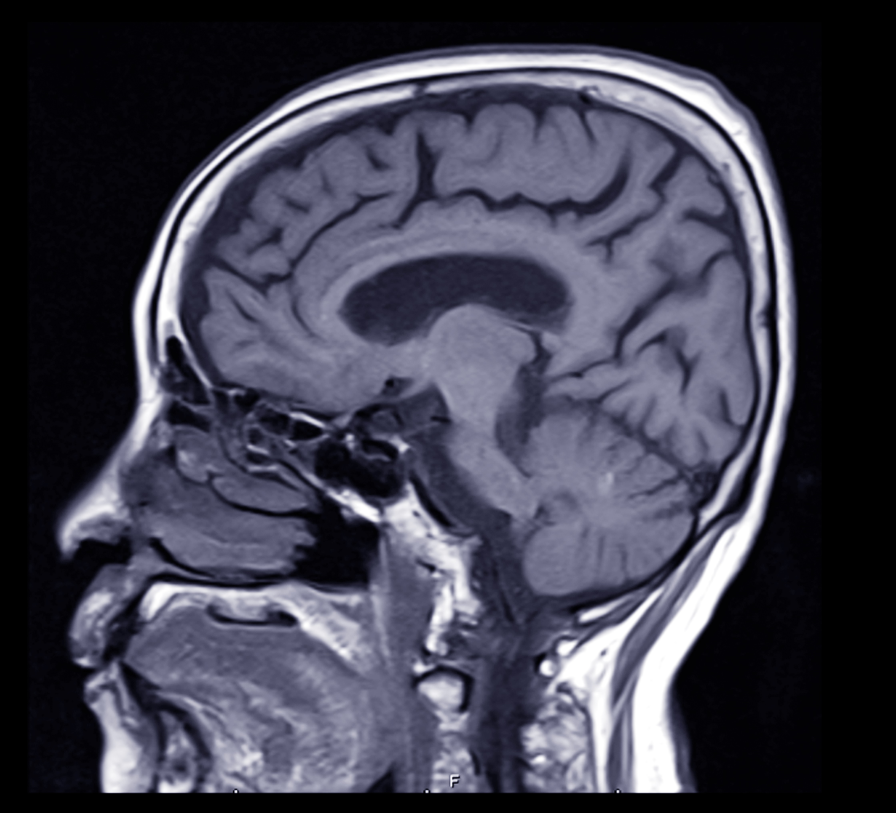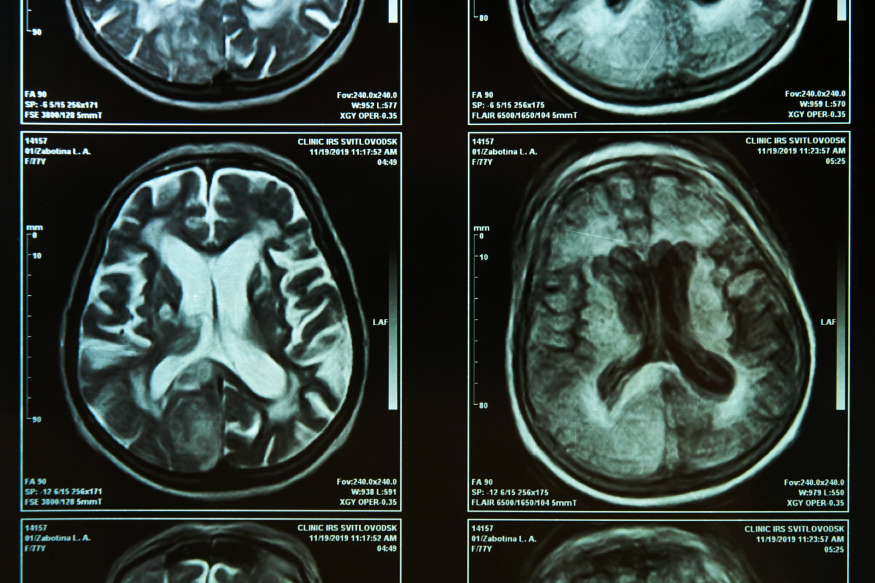
Doctors and researchers studying the molecular and clinical aspects of Alzheimer’s disease are learning more about the mechanisms of this devastating condition.
Published March 1, 2003
By Vida Foubister

While proteins involved in the generations of Alzheimer’s disease (AD) continue to perplex researchers, progress is being made in the presymptomatic and early identification of patients.
But the use of genetic testing, brain imaging and other available technologies to identify people who are likely to develop Alzheimer’s will remain problematic until disease modifying therapies become available, according to Norman Relkin, MD, PhD, associate professor of Clinical Neurology and Neuroscience at New York-Presbyterian Hospital, Weill Medical College of Cornell University.
“This information is viewed as toxic information that is potentially very, very harmful,” Relkin told participants at the Fourth New York Alzheimer’s Research Symposium held last Nov. 20. “It’s one thing to tell someone they’re at risk, it’s quite another thing for there to be nothing that they can do about it.”
The lifetime risk of AD is estimated to be 10 to 15 percent among the general population, meaning that one-in-10 women who live to 80 and one-in-seven men who live to 76 will develop the disease. This risk doubles to 20 to 30 percent for first-degree relatives – mother, father, sister, brother, daughter or son – of people with AD.
Convened by The New York Academy of Sciences (the Academy), the afternoon symposium brought together three experts on the evolving molecular and clinical aspects of Alzheimer’s disease. Its co-sponsors included The Institute for the Study of Aging, The New York City Chapter of the Alzheimer’s Association and The New York City Metro Area Chapter of the Society for Neuroscience.
The Calsenilin Story Evolves
One of the main pathological hallmarks of AD is the presence of amyloid plaques in patient brains. These plaques are composed of amyloid beta-peptide (Aβ), which is derived from gamma-secretase cleavage of the beta-amyloid precursor protein (APP).
Several years ago Joseph Buxbaum, PhD, associate professor of Psychiatry and head of the Laboratory of Molecular Neuropsychiatry at Mount Sinai School of Medicine, identified a novel protein that interacts with presenilin, a protein required for γ−secretase cleavage of APP that many scientists believe should be inhibited to prevent or treat Alzheimer’s. Named calsenilin, for calcium and presenilin binding protein, it was subsequently identified by other researchers as a transcription factor regulating dynorphin expression (DREAM) and a potassium channel interacting protein (KChIP3).
Buxbaum developed a calsenilin knockout mouse to determine the true physiological function of this protein. This work is important because it helps to identify the side effects of potential Alzheimer’s drugs, in this case drugs targeting presenilin that might affect calsenilin function as well. But calsenilin “has resisted, very effectively, easy analysis,” he said.
His initial results, which found Aβ formation and K+ currents decreased and long-term potentiation in the dentate gyrus of the hippocampus increased, suggested a role for calsenilin in regulating presenilin and voltage-gated potassium channel (Kv4) function. Further, the animals were found to be more sensitive to shock and, therefore, it seemed unlikely that calsenilin was involved in modulating pain sensitivity through the antagonism of dynorphin expression.
Calsenilin as a Dynorphin Suppressor
But data published by another lab showing that calsenilin knockouts were less sensitive to pain led Buxbaum to reevaluate the knockout using the tail-flick flick latency test. This test measures the time taken for a mouse to flick its tail away from a heat source, and has long been thought to be analogous with shock sensitivity.
The tail-flick results confirmed that the knockout mice were less sensitive to pain, and thus a role for calsenilin as a dynorphin suppressor could not be ruled out. These results also are likely to get much attention from those working in pain research. “The take-home message is that shock sensitivity is actually not reflective at all of tail-flick sensitivity,” he said.
Though the exact role of calsenilin in Alzheimer’s disease remains unclear, Buxbaum’s current hypothesis is that calsenilin affects Aβ formation by modulating calcium levels in the cell. There is a relationship between calcium abnormalities and Alzheimer’s, and it’s known that increases in cellular calcium result in the production of more and more Aβ.
Selective Degeneration
A pathological feature of AD, in addition to amyloid plaque and neurofibrillary tangle formation, is selective neurodegeneration. “In patients with Alzheimer’s disease, not all neurons are dying at the same time,” said Tae-Wan Kim, PhD, assistant professor at Columbia University’s Taub Institute for Research on Alzheimer’s Disease and the Aging Brain. “There is remarkable specific and selective degeneration and this underlies a lot of cognitive deficits.”
Focusing on the basal forebrain cholinergic neurons, a prime site of neuronal death in AD patients that also correlates with their cognitive deficits, Kim used proteomics to identify novel substrates that are cleaved by γ-secretase. He did this by comparing the protein profile of normal cells to those lacking functional γ-secretase.
“We wanted to find substrates expressed predominantly in these neurons that are affected in AD,” Kim explained. Notch, a developmentally regulated protein important for cell plate determination and neurogenesis, and ErbB4, a receptor tyrosine kinase, have previously been identified as γ-secretase substrates. Like APP, these substrates are cleaved in the transmembrane region and their cleavage is dependent on functional presenilins, early-onset familial AD genes.
Kim’s recent analysis found that the p75 neurotrophin receptor (p75-NTR), a protein that has been implicated in Alzheimer’s and other diseases due to its regulation of cell survival and death, is a γ-secretase substrate. He did this by demonstrating that p75-NTR undergoes ectodomain shedding, a step that is required for γ-secretase cleavage, and that the p75-NTR cleavage is blocked in cells treated with a γ-secretase inhibitor. Further, the p75-NTR cleavage site was found to be in the transmembrane region and similar to that of Aβ40, one of the APP peptide fragments.
Risk and Risk Perception
In future work, Kim plans to investigate whether shedding and cleavage by γ-secretase can regulate neuronal cell death and survival. “If that is the case then we might have a molecular basis for selective neurodegeneration of basal forebrain cholinergic neurons in AD,” he said. He also would like to use microarray analysis to identify downstream target genes.
Eighteen months before announcing he had been diagnosed with Alzheimer’s disease, former President Ronald Reagan gave a speech of about 90 minutes in length without a single error.
“He spoke perfectly,” said Relkin, who assessed a videotape of the 1992 speech for signs or symptoms of incipient AD. “When one considers that we’re trying to come to a point in which we can diagnose AD in its presymptomatic stages, or at least predict with reasonable accuracy who is going to develop the disease, performances like that are daunting.”
Relkin also sees it as a lesson that predictive testing should involve more than clinical interviews and observational methods.
The field has moved from diagnosing Alzheimer’s by exclusion to direct diagnosis. In addition, clinicians have begun subcategorizing patients with mild cognitive impairment (MCI) who are believed to have an increased likelihood of developing AD or other forms of dementia in the near future.
Managing Risk Perception
One of these subgroups, AD-like MCI, includes patients whose symptoms are found to have an “AD-like flavor.” It’s estimated that from 5 to 40 percent of them go on to develop AD each year.
Translating this subcategorization into general practice without more specific diagnostic criteria, however, will be problematic. This is where technologies such as genetic testing, proteomic analysis and structural/functional neuroimaging can be used to improve differential diagnosis and presymptomatic detection of AD.
As more information becomes available, managing patients’ risk perception will become important. The general population is more influenced by their perceptions of risk than by numbers representing the probability they will develop a disease, said Relkin. “Perceptions are altered by life experiences, like caring for patients in the family with the disease, and have a greater impact on how one views one’s risk of AD.”
Results he presented from an ongoing study, called REVEAL (Risk Evaluation and Education for Alzheimer’s Disease), confirm this. People in the survey tended to remember their genotype, that is whether or not they carry the APOE 4 allele associated with an increased risk of AD, more than their numeric lifetime risk estimates.